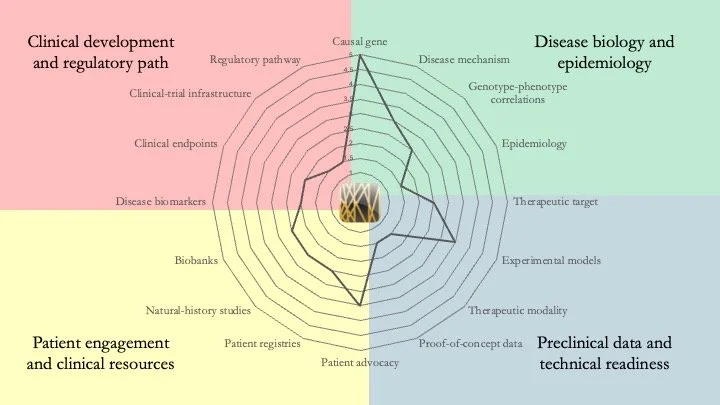
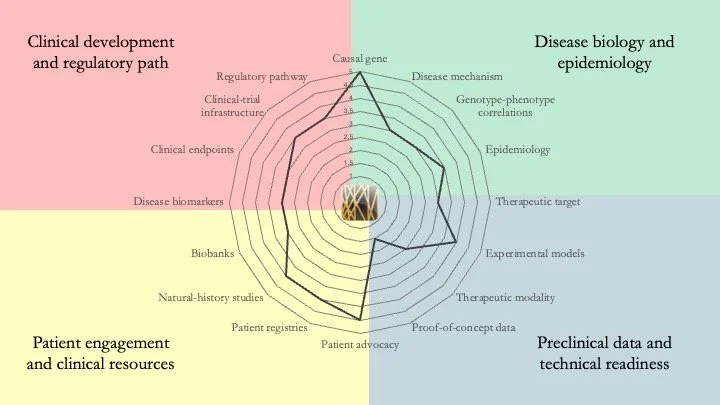
Our research-readiness assessments provide an at-a-glance view of the gaps that should be addressed before and during the execution of a drug-discovery project. We also create bespoke reports that provide a detailed analysis of the readiness of your ultra-rare disease of interest, helping you sharpen your research strategy and prioritize your goals. Get in touch to find out how we can help your organization.
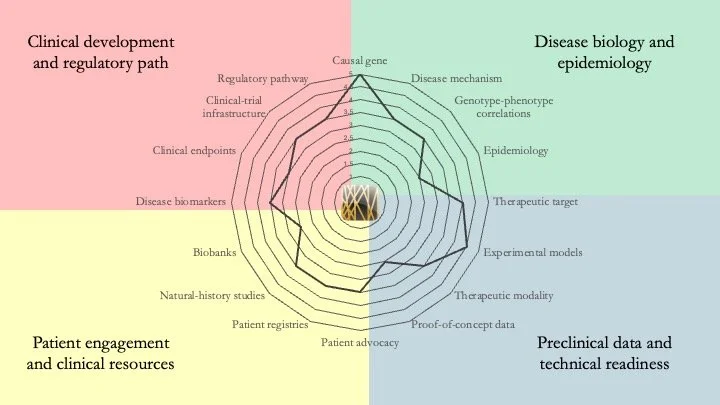
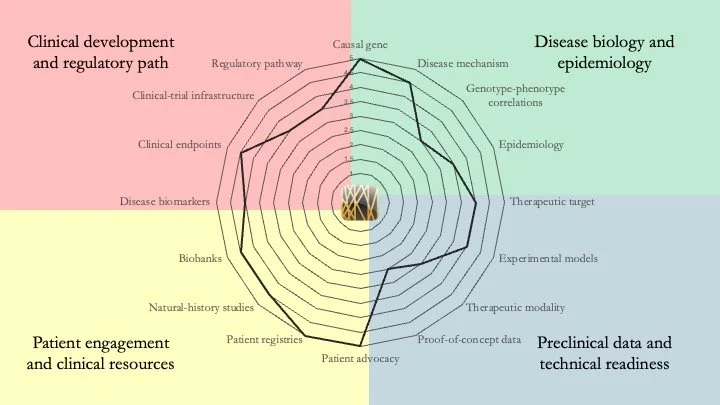
ADNP Syndrome. Also known as Helsmoortel–Van der Aa Syndrome, this is a neurodevelopmental disorder caused by mutations in the gene encoding Activity-Dependent Neuroprotective Protein. Our assessment shows that this condition is sufficiently mature for clinical development. In fact, clinical programs have already been launched. However, gaps remain in terms of mechanistic understanding and clinical resources. April 30, 2026.
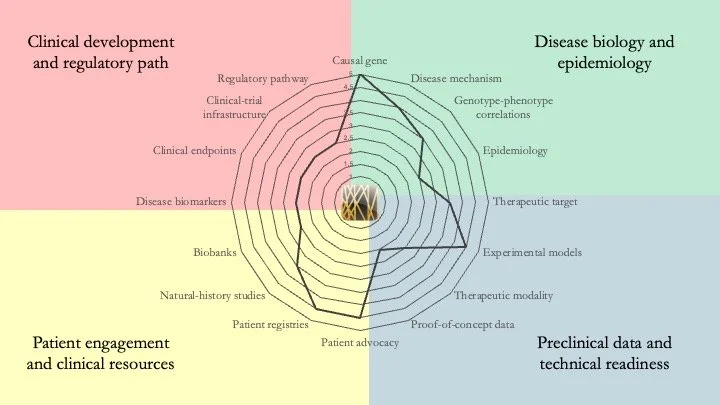
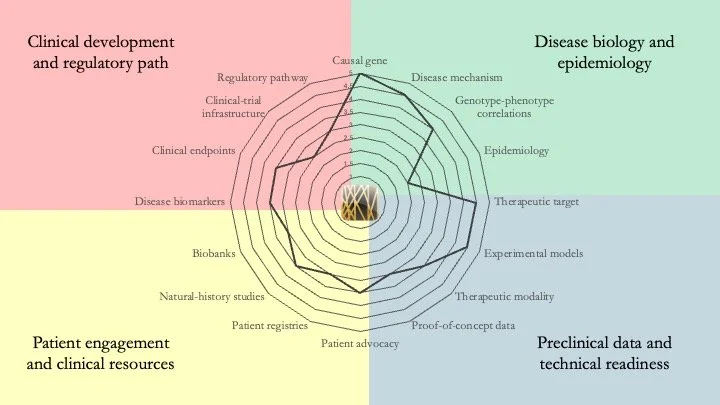
DYRK1A Syndrome. DYRK1A haploinsufficiency results in developmental delay, intellectual disability and microcephaly. There is a lot of preclinical work around this condition, and a recent study by Ron et al. in the journal Molecular Psychiatry on the therapeutic effect of lithium on a mouse model stands out as particularly important. April 27, 2026.
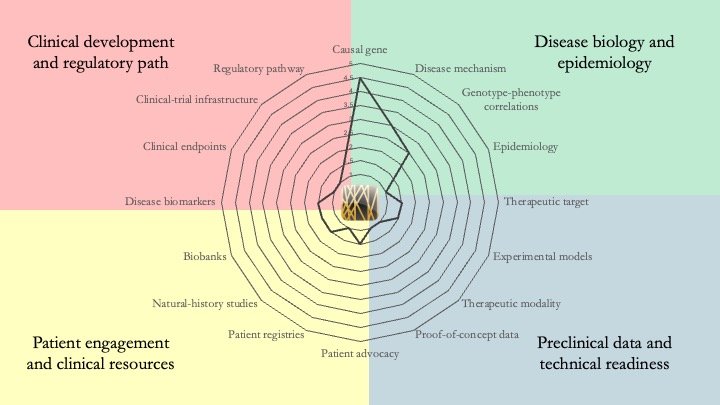
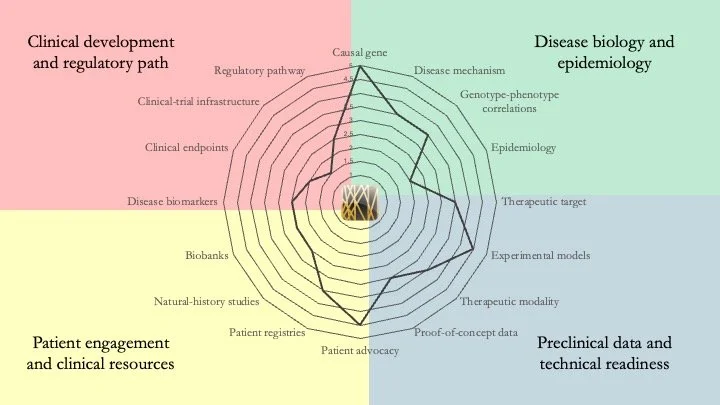
TFE3-associated neurodevelopmental disorder. Only a few dozen patients have been identified since the disease was first described in 2019, which explains why multiple readiness gaps remain to be addressed. But what makes the condition particularly interesting is that the underlying mechanism may be amenable to the use of molecular glues, something that isn't always the case for genetic diseases. April 23, 2026.
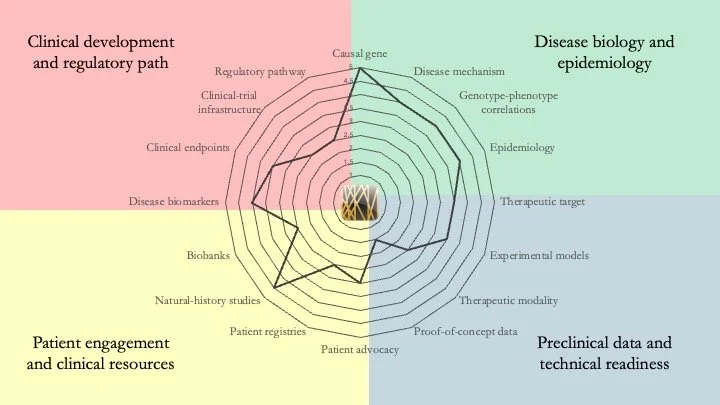
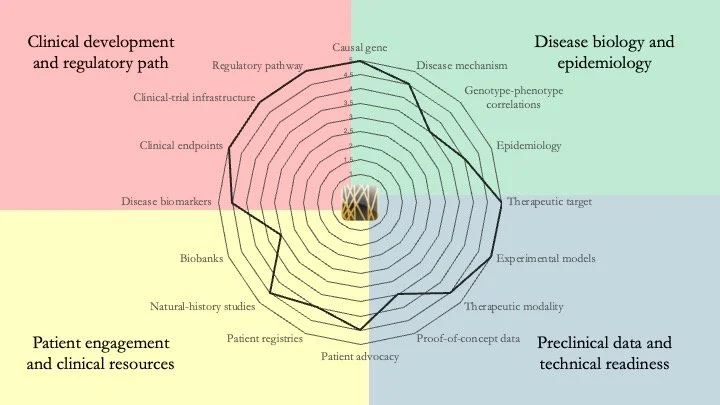
HRPT1 deficiency. HRPT1 encodes hypoxanthine-guanine phosphoribosyltransferase, a key enzyme in the purine salvage pathway. Its deficiency results in a variety of diseases, some of which are very severe (e.g. Lesch-Nyhan Disease). Our assessment indicates that phenotype-genotype correlations are well established for this group of diseases, but the absence of robust animal models has prevented translational progress. April 16, 2026.
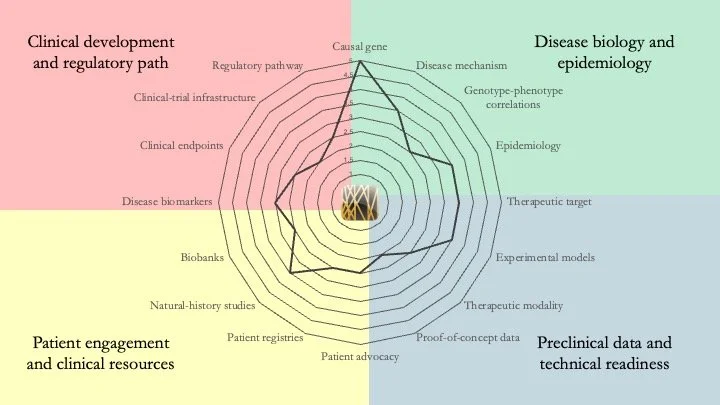
Glycogen Storage Disease IV. Also known as Andersen Disease, this condition is cause by mutations in GBE1, which encodes 1,4-alpha-glucan branching enzyme 1. Broadly speaking, there are two forms of the disease: a pediatric form and an adult-onset form. The research readiness of the two forms is different, and the above plot shows a composite score, which discloses multiple gaps in different domains. April 13, 2026.
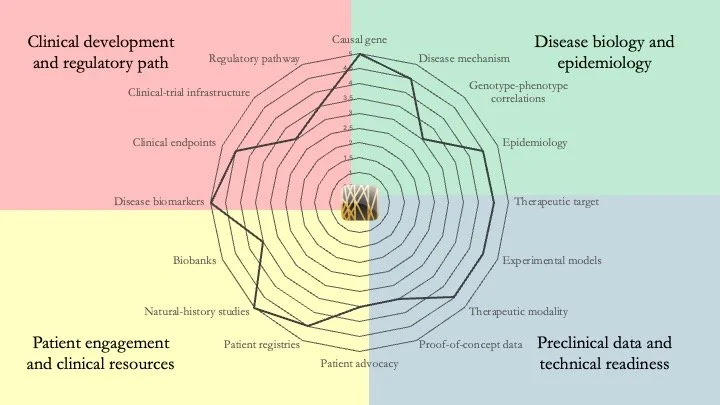
Glutaric Aciduria Type 1. This disease is an inborn error of metabolism caused by mutations in the GCDH gene, preventing the breakdown of lysine, hydroxylysine and tryptophan. It is accompanied by metabolic crises and neurological damage. Fortunately, our understanding of the condition is quite mature and on the cusp of clinical translation. April 9, 2026.
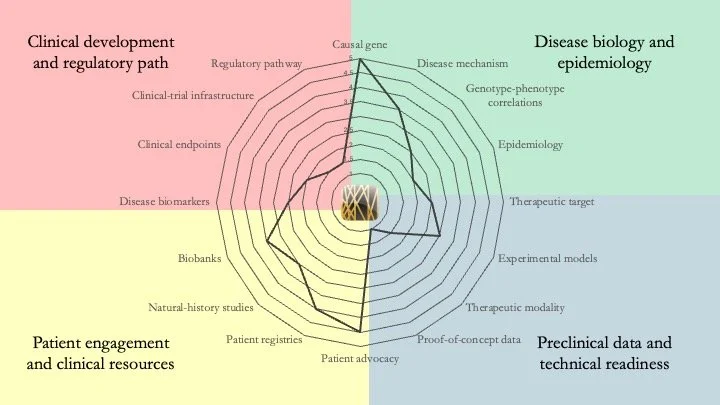
Bohring-Opitz Syndrome. This is a multi-system disorder caused by mutations in ASXL1, which codes for a protein involved in chromatin remodeling. Our analysis shows that a lot of work remains to be done in terms of technical readiness and laying the foundation for eventual clinical development. An immediate priority would be to identify the right therapeutic modality to use and the right phenotype to target. April 6, 2026.
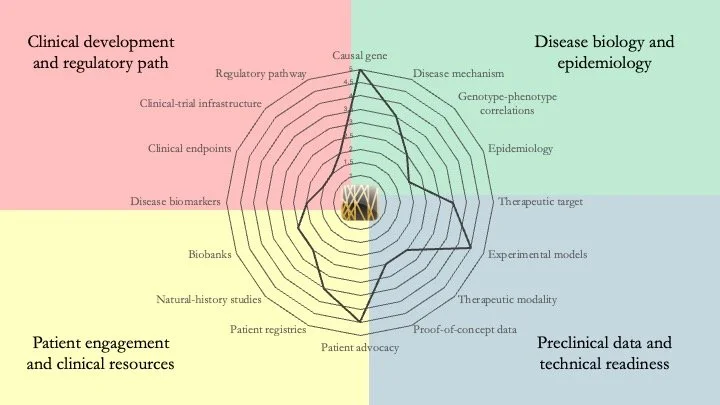
MEF2C Haploinsufficiency Syndrome. This condition, caused by mutations in one copy of the transcription factor MEF2C, results in severe developmental delay, epilepsy and sleep disturbances, among other symptoms. Our research-readiness assessment shows that, although the disease mechanism is increasingly well understood, key gaps include the paucity of rescue data in preclinical models using any disease-modifying therapeutic modality and the absence of clinical-trial infrastructure. April 2, 2026.
Shwachman-Diamond Syndrome. This multi-system disorder is caused by mutations in the SBDS gene, a protein important for ribosomal biogenesis. The disease is characterized by bone-marrow failure, pancreatic insufficiency, skeletal abnormalities and other phenotypes. The main gaps that we identified pertain to the lack of in vivo data showing phenotypic rescue after therapeutic interventions. The emergence of new experimental models may help address this issue. March 30, 2026.
Timothy Syndrome. This channelopathy is characterized by the presence of arrhythmias and neurodevelopmental phenotypes. Timothy Syndrome is caused by gain-of-function mutations in CACNA1C, which encodes the voltage-gated calcium channel Cav1.2. Our analysis shows that proof-of-concept data remain scarce, with ASOs being the preferred modality so far. Key gaps also remain in terms of clinical readiness and establishing the prevalence of the syndrome. March 26, 2026.
PACS1-related syndrome. Also known as Schuurs-Hoeijmakers Syndrome, this neurodevelopmental disorder is caused by mutations in a protein involved in trafficking through the Golgi network. The existence of strong genotype-phenotype correlations point to ASOs as the ideal therapeutic modality for this disease. However, important gaps remain on the clinical-development side. March 23, 2026.
Gorlin Syndrome. Condition caused by mutations in PTCH1 and characterized by the development of basal cell carcinomas and other tumors, as well as skeletal abnormalities. Our analysis shows very few gaps, which is not unexpected, considering the availability of therapies for this condition. But as the existing therapies are not curative, Gorlin Syndrome may be fertile ground for the development of advanced therapeutics. March 19, 2026.
ADSSL1 Myopathy. Mutations in the ADSSL1 gene, which encodes adenylosuccinate synthetase 1, affect purine metabolism and energy production in muscle cells. Our analysis shows that multiple gaps remain across the different domains that we evaluate, starting with a limited understanding of disease biology. March 16, 2026.
Alström Syndrome. Multi-system disorder caused by mutations in the ALMS1 gene. Our analysis indicates that important gaps remain in understanding disease mechanisms and in the identification of the ideal modality to develop a therapeutic. March 12, 2026.